A brief foray into parallel processing with R
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
I’ve recently been dabbling with parallel processing in R and have found the foreach package to be a useful approach to increasing efficiency of loops. To date, I haven’t had much of a need for these tools but I’ve started working with large datasets that can be cumbersome to manage. My first introduction to parallel processing was somewhat intimidating since I am surprisingly naive about basic computer jargon – processors, CPUs, RAM, flux capacitors, etc. According the the CRAN task view, parallel processing became directly available in R beginning with version 2.14.0, and a quick look at the web page provides an astounding group of packages that explicitly or implicitly allow parallel processing utilities.
In my early days of programming I made liberal use of for loops for repetitive tasks. Not until much later I realized that for loops are incredibly inefficient at processing data. This is common knowledge among programmers but I was completely unaware of these issues given my background in the environmental sciences. I had always assumed that my hardware was more than sufficient for any data analysis needs, regardless of poor programming techniques. After a few watershed moments I soon learned the error of my ways and starting adopting more efficient coding techniques, e.g., vectorizing, apply functions, etc., in addition to parallel processing.
A couple months ago I started using the foreach package with for loops. To be honest, I think loops are unavoidable at times regardless of how efficient you are with programming. Two things struck me when I starting using this package. First, I probably could have finished my dissertation about a year earlier had I been using parallel processing. And two, the functions are incredibly easy to use even if you don’t understand all of the nuances and jargon of computer speak. My intent of this blog is to describe how the foreach package can be used to quickly transform traditional for loops to allow parallel processing. Needless to mention, numerous tutorials covering this topic can be found with a quick Google search. I hope my contribution helps those with little or no experience in parallel processing to adopt some of these incredibly useful tools.
I’ll use a trivial example of a for loop to illustrate repeated execution of a simple task. For 10 iterations, we are creating a normally-distributed random variable (1000000 samples), taking a summary, and appending the output to a list.
#number of iterations in the loop
iters<-10
#vector for appending output
ls<-vector('list',length=iters)
#start time
strt<-Sys.time()
#loop
for(i in 1:iters){
#counter
cat(i,'\n')
to.ls<-rnorm(1e6)
to.ls<-summary(to.ls)
#export
ls[[i]]<-to.ls
}
#end time
print(Sys.time()-strt)
# Time difference of 2.944168 secs
The code executes quickly so we don’t need to worry about computation time in this example. For fun, we can see how computation time increases if we increase the number of iterations. I’ve repeated the above code with an increasing number of iterations, 10 to 100 at intervals of 10.
#iterations to time
iters<-seq(10,100,by=10)
#output time vector for iteration sets
times<-numeric(length(iters))
#loop over iteration sets
for(val in 1:length(iters)){
cat(val,' of ', length(iters),'\n')
to.iter<-iters[val]
#vector for appending output
ls<-vector('list',length=to.iter)
#start time
strt<-Sys.time()
#same for loop as before
for(i in 1:to.iter){
cat(i,'\n')
to.ls<-rnorm(1e6)
to.ls<-summary(to.ls)
#export
ls[[i]]<-to.ls
}
#end time
times[val]<-Sys.time()-strt
}
#plot the times
library(ggplot2)
to.plo<-data.frame(iters,times)
ggplot(to.plo,aes(x=iters,y=times)) +
geom_point() +
geom_smooth() +
theme_bw() +
scale_x_continuous('No. of loop iterations') +
scale_y_continuous ('Time in seconds')
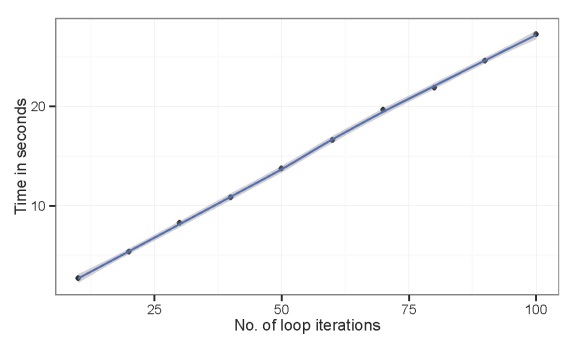
The processing time increases linearly with the number of iterations. Again, processing time is not extensive for the above example. Suppose we wanted to run the example with ten thousand iterations. We can predict how long that would take based on the linear relationship between time and iterations.
#predict times mod<-lm(times~iters) predict(mod,newdata=data.frame(iters=1e4))/60 # 45.75964
This is all well and good if we want to wait around for 45 minutes. Running the loop in parallel would greatly decrease this time. I want to first illustrate the problem of running loops in sequence before I show how this can done using the foreach package. If the above code is run with 1e4 iterations, a quick look at the performance metrics in the task manager (Windows 7 OS) gives you an idea of how hard your computer is working to process the code. My machine has eight processors and you can see that only a fraction of them are working while the script is running.

for loop.
Running the code using foreach will make full use of the computer’s processors. Individual chunks of the loop are sent to each processor so that the entire process can be run in parallel rather than in sequence. That is, each processor gets a finite set of the total number of iterations, i.e., iterations 1–100 goes to processor one, iterations 101–200 go to processor two, etc. The output from each processor is then compiled after the iterations are completed. Here’s how to run the code with 1e4 iterations in parallel.
#import packages
library(foreach)
library(doParallel)
#number of iterations
iters<-1e4
#setup parallel backend to use 8 processors
cl<-makeCluster(8)
registerDoParallel(cl)
#start time
strt<-Sys.time()
#loop
ls<-foreach(icount(iters)) %dopar% {
to.ls<-rnorm(1e6)
to.ls<-summary(to.ls)
to.ls
}
print(Sys.time()-strt)
stopCluster(cl)
#Time difference of 10.00242 mins
Running the loop in parallel decreased the processing time about four-fold. Although the loop generally looks the same as the sequential version, several parts of the code have changed. First, we are using the foreach function rather than for to define our loop. The syntax for specifying the iterator is slightly different with foreach as well, i.e., icount(iters) tells the function to repeat the loop a given number of times based on the value assigned to iters. Additionally, the convention %dopar% specifies that the code is to be processed in parallel if a backend has been registered (using %do% will run the loop sequentially). The functions makeParallel and registerDoParallel from the doParallel package are used to create the parallel backend. Another important issue is the method for recombining the data after the chunks are processed. By default, foreach will append the output to a list which we’ve saved to an object. The default method for recombining output can be changed using the .combine argument. Also be aware that packages used in the evaluated expression must be included with the .packages argument.
The processors should be working at full capacity if the the loop is executed properly. Note the difference here compared to the first loop that was run in sequence.

for loop.
A few other issues are worth noting when using the foreach package. These are mainly issues I’ve encountered and I’m sure others could contribute to this list. The foreach package does not work with all types of loops. I can’t say for certain the exact type of data that works best, but I have found that functions that take a long time when run individually are generally handled very well. For example, I chose the above example to use a large number (1e6) of observations with the rnorm function. Interestingly, decreasing the number of observations and increasing the number of iterations may cause the processors to not run at maximum efficiency (try rnorm(100) with 1e5 iterations). I also haven’t had much success running repeated models in parallel. The functions work but the processors never seem to reach max efficiency. The system statistics should cue you off as to whether or not the functions are working.
I also find it bothersome that monitoring progress seems is an issue with parallel loops. A simple call using cat to return the iteration in the console does not work with parallel loops. The most practical solution I’ve found is described here, which involves exporting information to a separate file that tells you how far the loop has progressed. Also, be very aware of your RAM when running processes in parallel. I’ve found that it’s incredibly easy to max out the memory, which not only causes the function to stop working correctly, but also makes your computer run like garbage. Finally, I’m a little concerned that I might be destroying my processors by running them at maximum capacity. The fan always runs at full blast leading me to believe that critical meltdown is imminent. I’d be pleased to know if this is an issue or not.
That’s it for now. I have to give credit to this tutorial for a lot of the information in this post. There are many, many other approaches to parallel processing in R and I hope this post has been useful for describing a few of these simple tools.
Cheers,
Marcus
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
